A. Describe the characteristics a cloning vector must possess.
B. Why DNA cannot pass through the cell membrane? Explain. How is a bacterial cell made competent to take up recombinant DNA from the medium?
A. Characteristics features of cloning vector
1. Presence of Origin of replication (ori) - The vector should have an ori which is a sequence from where replication starts. Any piece of DNA which is linked to the ori can be made to replicate within the host cells. The ori site also controls the copy number so cloning vectors having ori that support a high copy number are chosen.
2. Selectable marker – The vector should possess a selectable marker which helps to distinguish and identify the non-transformants from transformants and selectively permit the growth of transformants. Genes encoding for antibiotics like ampicillin, tetracycline, chloramphenicol or kanamycin are considered to be useful selectable markers for E.coli.
3. Cloning Sites – The cloning vector should contain a single recognition site for the restriction enzymes in order to link the alien DNA. The presence of more than one recognition sites generates several fragments and complicates the cloning process.
4. It should have a high copy number so that we obtain many copies of the DNA linked to it
5. They should be able to replicate independently.
6. They should help easy linking of alien DNA.
B. DNA cannot pass through the cell membranes as it is a hydrophilic molecule. The cell membrane is made up of lipid bilayer which makes it difficult for the hydrophilic DNA molecule to pass through it.
Bacterial cells are made competent to take up recombinant DNA from the medium in the following ways-:
1. Treating bacteria with specific concentration of divalent cation such as calcium that results in an increase in the efficiency with which DNA is taken up by the bacterial cell. Recombinant DNA can then be forced into such cells by incubating them with the recombinant DNA on ice followed by placing them briefly at 42°C (heat shock), and again back on the ice.
Some other methods available for introduction of desired DNA are:
1. Microinjection in which the recombinant DNA is directly injected into the nucleus of an animal cell.
2. Biolistics or gene gun method is suitable for plant cells, where cells are bombarded with high-speed micro-particles of gold or tungsten coated with DNA.
If a desired gene is identified in an organism for some experiments, explain the process of the following:
(i) Cutting this desired gene at specific location
(ii) Synthesis of multiple copies of this desired gene
I. Restriction enzymes - Cutting of the desired gene at a specific location is attained by using restriction endonucleases. These enzymes are specialised to cut the fragment of DNA at specific locations. Each restriction endonuclease functions by ‘inspecting’ the length of a DNA sequence. Once it finds its specific recognition sequence, it binds to the DNA and cuts each of the two strands of the double helix at specific points in their sugar -phosphate backbones. Each restriction endonuclease recognises a specific palindromic nucleotide sequence (sequence of base pairs that reads same on the two strands when the orientation of reading is kept the same) in the DNA. Restriction enzymes cut the strand of DNA a little away from the centre of the palindrome sites, but between the same two bases on the opposite strands. This leaves single-stranded portions at the ends.
II. Polymerase Chain Reaction - PCR is used to create multiple copies of the DNA being incorporated with molecular biology tools to obtain a higher copy of the desired gene. Two sets of chemically synthesised oligonucleotides and DNA polymerase are being used in vitro for the multiplication of the desired gene.
The PCR produces approximately billion copies of a gene in less than 20 minutes. Such higher number of the product is achieved by use of thermostable DNA polymerase (isolated from a bacterium, Thermus aquaticus.
Components used during PCR - Template DNA, DNA polymerase, Primers and buffer.
Steps of PCR – •
Denaturation - The DNA is denatured that is the strands are separated by heating the dsDNA to 95-degree celsius. This breaks hydrogen bonds that hold DNA strands together in the helix. •
Annealing- The mixture is cooled, this allows the primer to bind to their respective complementary sequence. •
Extension- The mixture is then heated so that DNA polymerase can synthesise new strands.
The whole cycle is repeated to create multiple copies.
Which of the following is a restriction endonuclease?
Protease
DNase I
RNase
RNase
D.
RNase
Hind II is a restriction endonuclease are enzymes used for cutting DNA at specific locations.
(a) Describe the various steps of Griffith’s experiment that led to the conclusion of the ‘Transforming Principle’.
(b) How did the chemical nature of the ‘Transforming Principle’ get established?
Grifith's experiment
i. Streptococcus pneumoniae (pneumococcus) bacteria were grown on a culture plate, some produces smooth shiny colonies (S) while others produces rough colonies (R). This was because the S strain bacteria has a mucous (polysaccharide) coat, while R strain does not.
Mice infected with the S strain (virulent) die from pneumonia infection but mice infected with the R strain do not develop pneumonia.
S strain ---> Inject into mice ----> Mice die
R strain ----> Inject into mice ----> Mice live
The virulet S strain bacteria were killed by heating them. It was observed that heat killed S strain bacteria injected into mice did not kill them.
S strain (heat-killed) ----> Inject into mice ----> Mice live
When a mixture of heat-killed S and live R bacteria was injected, the mice died.
S strain (heat-killed) + R strain (live) ----> Inject into mice ----> Mice died. Moreover, living S bacteria was recovered from the dead mice.
The experiment concluded that the R strain bacteria had somehow been transformed by the heat-killed S strain bacteria and that some ‘transforming principle’, transferred from the heat-killed S strain, had enabled the R strain to become virulent. It was thought that this must be due to the transfer of the genetic material.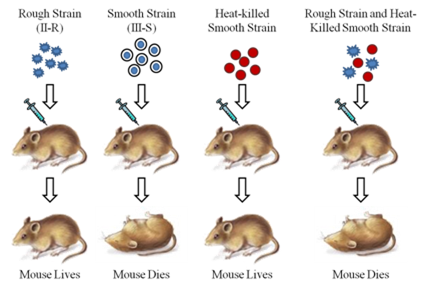
b. Prior to the work of Oswald Avery, Colin MacLeod and Maclyn McCarty it was thought that the genetic material was a protein. They worked to determine the biochemical nature of ‘transforming principle’ in Griffith’s experiment. They purified biochemicals (proteins, DNA, RNA, etc.) from the heat-killed S cells to see which ones could transform live R cells into S cells. They discovered that protein-digesting enzymes (proteases) and RNA digesting enzymes (RNases) did not affect transformation, so the transforming substance was not a protein or RNA. Digestion with DNase did inhibit transformation, suggesting that the DNA caused the transformation. They concluded that DNA is the hereditary material and that DNA alone from S bacteria caused R bacteria to become transformed.
